|
7/26/2023 0 Comments Okazaki fragment sequencing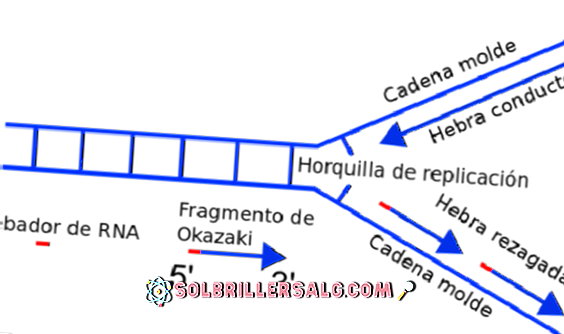
The next round of replication results in two daughter cells with the rearrangements depicted in A. The top strand resembles the bottom and original strain in A, and the bottom strand resembles the top and mutated strain in A. (D) The synthesized strand aligns back to the chromosome at point 4 (template switch #2) and (E) continues synthesis in the original direction. (C) At point 2, the DNA folds back (template switch #1) and synthesizes until it arrives at point 3. (B) DNA synthesis starts on a template similar to the bottom line in A and continues until the fork is stalled (point 1). Numbers indicate important points in the mechanism. Arrows indicate palindromes, and brackets indicate duplications. (A) The top line includes the mutated strain with an insertion mediated by a palindrome the bottom line is the original sequence. Palindrome expansion occurs via template switching in rad27Δ strains. The red strand is the nascent strand and the blue strand is the original template strand. Arrows indicate palindromes, and bases showing the differences between arms are shown in large font. The next round of replication results in two daughter cells that resemble the variants depicted in A. Finally, (D) the nascent strand returns to the correct template. A template switch occurs, causing the nascent strand to use itself as a template, eliminating variation between the arms of the palindrome. Then, the newly synthesized strand folds back using the homology between the arms. The fork synthesizes one arm and starts synthesis on the second arm (C). (B) Replication starts on a template similar to the bottom strain in A. The top sequence represents the mutated strain that includes a perfect palindrome, and the bottom sequence represents the original (nonmutated) strain that includes a quasi-palindrome. (A) Variation between two rad27Δ strains from subculture 25. 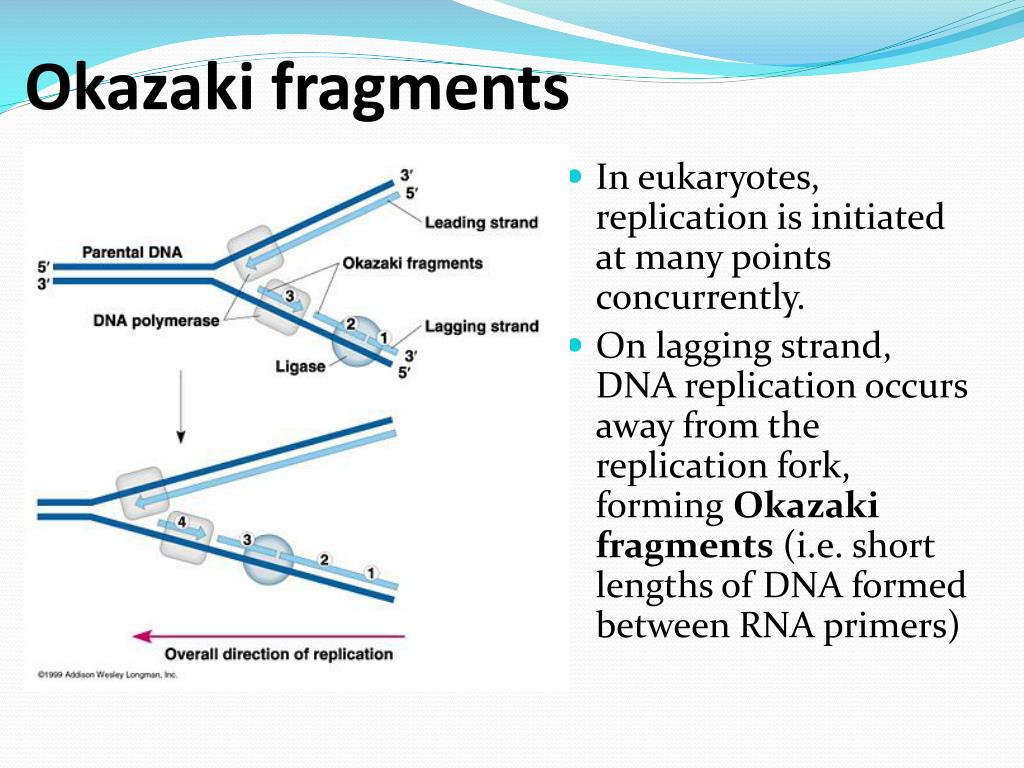 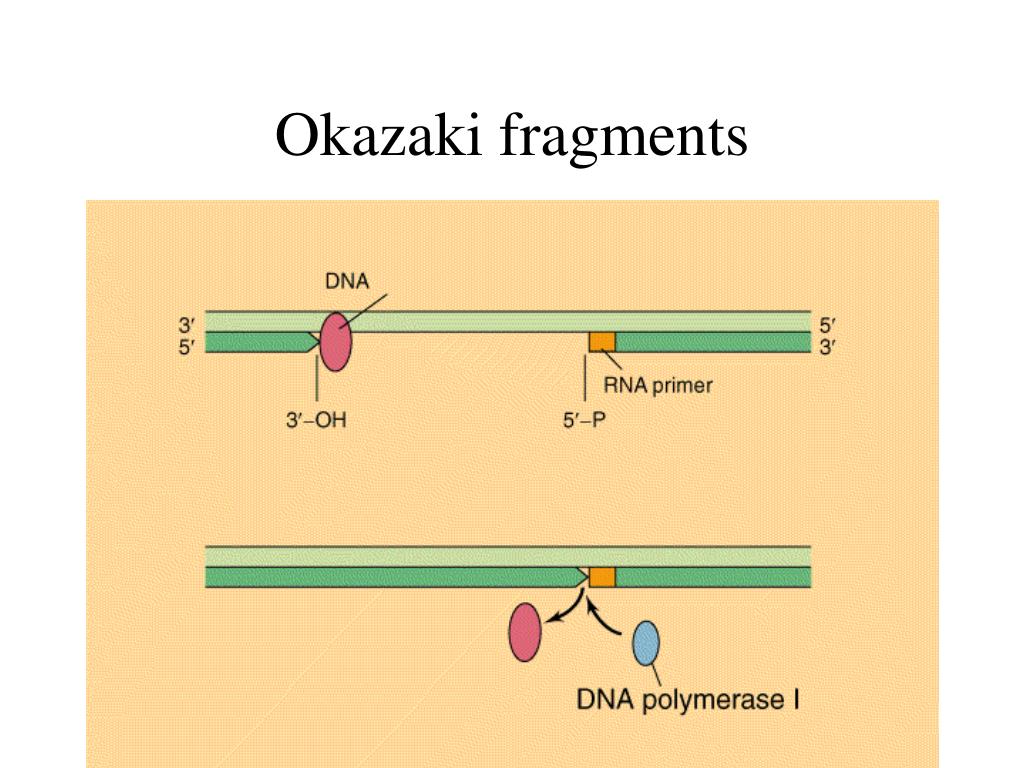
Quasi-palindrome to palindrome correction in rad27Δ strains. We note that our results open new ways of understanding template switching that occurs during genome instability and evolution.įEN1 Okazaki fragments RAD27 inverted repeats template switching. Since Rad27 acts on the lagging strand when the leading strand should not contain any gaps, we propose a mechanism favoring intramolecular strand switching over an intermolecular mechanism. Out of the 455 events, 55 events appeared in multiple isolates further analysis indicates that these loci are mutational hotspots. Evidence of quasi-palindrome to palindrome correction that could be generated by template switching appears also in yeast genome evolution. The formation of these events is best explained by folding back of the stalled nascent strand and resumption of DNA synthesis using the same nascent strand as a template. These events were presumably generated by quasi-palindrome to palindrome correction, as well as palindrome elongation. Surprisingly, we also detected a previously neglected class of 21 template-switching events. Out of the 455 changes observed in 10 colonies isolated the two most common types of events were insertions or deletions (INDELs) in simple sequence repeats (SSRs) and INDELs mediated by short direct repeats. cerevisiae rad27Δ strain was subcultured for 25 generations and sequenced using Illumina paired-end sequencing. To obtain a genomic view of the mutational consequence of loss of RAD27, a S. In Saccharomyces cerevisiae, a key player in this process is the structure-specific flap endonuclease, Rad27p (human homolog FEN1). The RNA fragment must be removed before the fragments are joined. Okazaki fragments that are formed during lagging strand DNA synthesis include an initiating primer consisting of both RNA and DNA.
0 Comments
Leave a Reply. |

 RSS Feed
RSS Feed
